Hi, I’m Xiongzhi Wang(王雄智, born in September 1995). I am currently pursuing my Ph.D. degree (combined Master–Ph.D. program) at the School of Aerospace Science and Technology, Xidian University, Xi’an, China.
I have also studied as an exchange student at the School of Future Technology, University of Chinese Academy of Sciences in Beijing,
and the Media Computing Lab at the College of Computer Science, Nankai University in Tianjin.
My research focuses on real-time stereo depth estimation, 3D reconstruction, and neural rendering. I specialize in developing high-performance perception frameworks for challenging environments, extending from minimally invasive surgery to autonomous driving and industrial digital twins. By combining mathematical modeling with deep learning, I strive to solve real-world perception bottlenecks through efficient, edge-deployable solutions.
🔥 News
- 2026.03: 🎉🎉 May God bless me.
📝 Publications
中科院2区
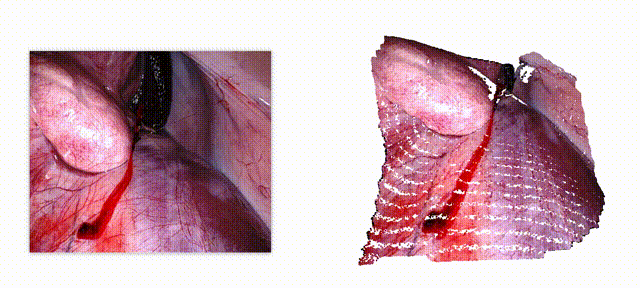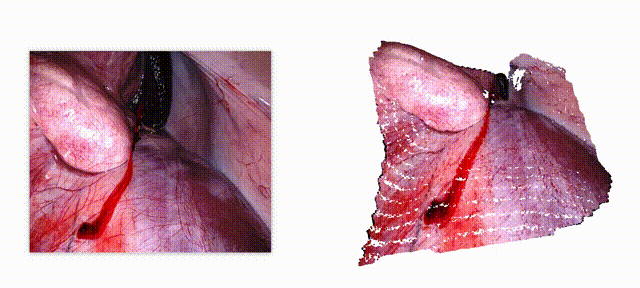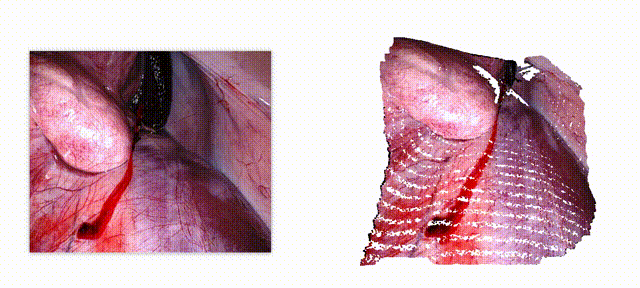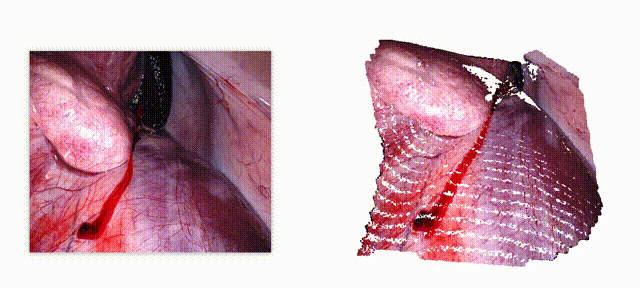
Endo-E2E-GS: End-to-end 3D reconstruction of endoscopic scenes using Gaussian Splatting
Xiongzhi Wang, Boyu Yang, Min Wei, Yu Chen, Jingang Zhang, Yunfeng Nie
Displays, 2026: 103353
- Merges depth estimation and Gaussian splatting for photometric supervision.
- Achieved superior quality on ENDONERF and SCARED surgical datasets.
- Enhances geometric perception for robotic-assisted endoscopic surgery workflows.
[paper]
[code]
[abstract]
Three-dimensional (3D) reconstruction is essential for enhancing spatial perception and geometric understanding in minimally invasive surgery. However, current methods like Neural Radiance Fields (NeRF) and 3D Gaussian Splatting (3DGS) often rely on offline preprocessing—such as COLMAP-based point clouds or multi-frame fusion—limiting their adaptability and clinical deployment. We propose Endo-E2E-GS, a fully end-to-end framework that reconstructs structured 3D Gaussian fields directly from a single stereo endoscopic image pair. The system integrates (1) a DilatedResNet-based stereo depth estimator for robust geometry inference in low-texture scenes, (2) a Gaussian attribute predictor that infers per-pixel rotation, scale, and opacity, and (3) a differentiable splatting renderer for 2D view supervision. Evaluated on the ENDONERF and SCARED datasets, Endo-E2E-GS achieves highly competitive performance, reaching PSNR values of 38.874/33.052 and SSIM scores of 0.978/0.863, respectively, surpassing recent state-of-the-art approaches. It requires no explicit scene initialization and demonstrates consistent performance across two representative endoscopic datasets.
中科院2区 Top
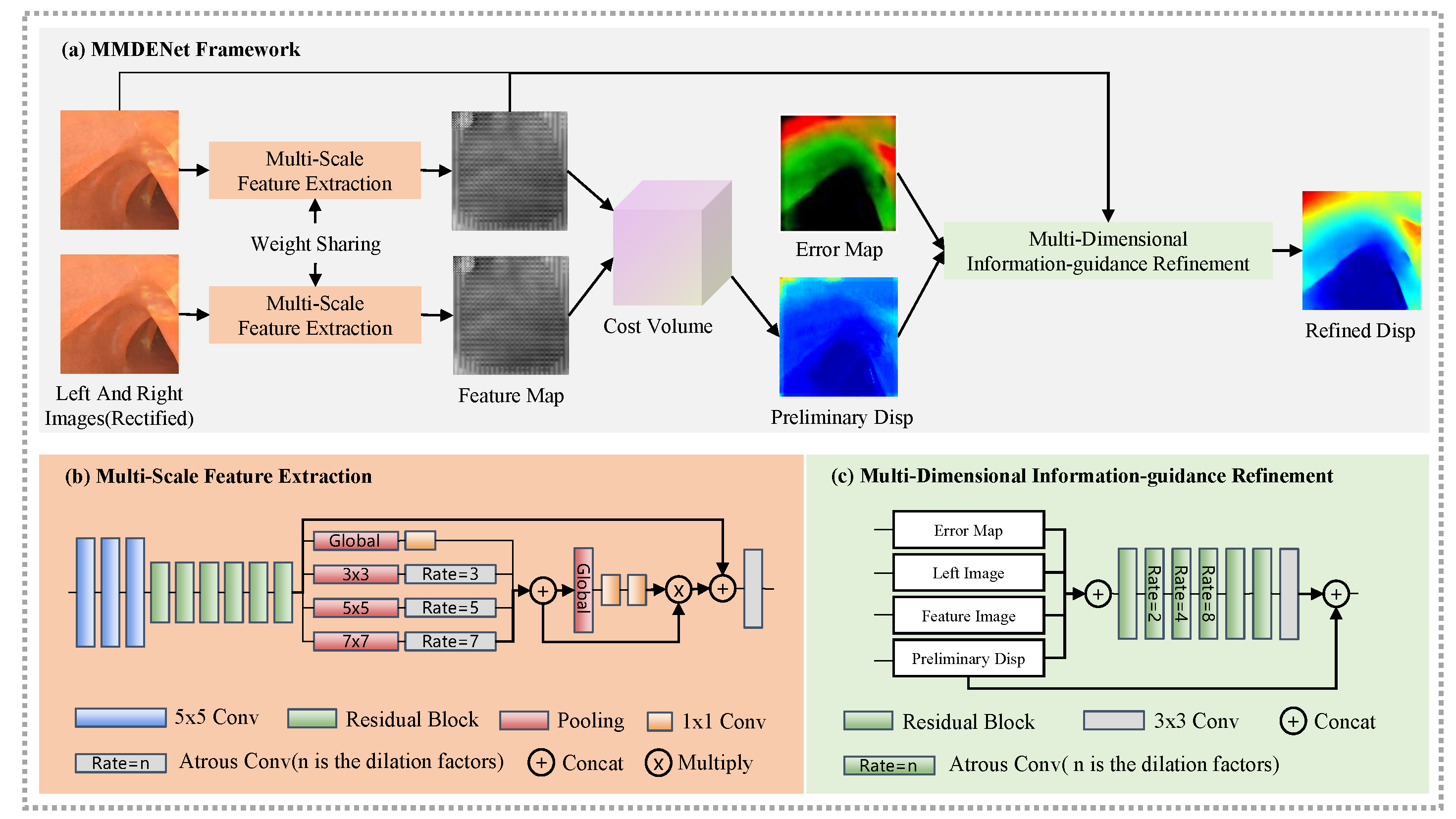
Multi-scale, multi-dimensional binocular endoscopic image depth estimation network
Xiongzhi Wang, Yunfeng Nie, Wenqi Ren, Min Wei, Jingang Zhang
Computers in Biology and Medicine, 2023, 164: 107305
- The depth estimation problem in the endoscopy environment is investigated.
- Method for generating endoscopic image datasets with depth information.
- A CNN network is proposed for estimating the depth of binocular image pairs.
[paper]
[abstract]
During invasive surgery, the use of deep learning techniques to acquire depth information from lesion sites in real-time is hindered by the lack of endoscopic environmental datasets. This work aims to develop a high-accuracy three-dimensional (3D) simulation model for generating image datasets and acquiring depth information in real-time. Here, we proposed an end-to-end multi-scale supervisory depth estimation network (MMDENet) model for the depth estimation of pairs of binocular images. The proposed MMDENet highlights a multi-scale feature extraction module incorporating contextual information to enhance the correspondence precision of poorly exposed regions. A multi-dimensional information-guidance refinement module is also proposed to refine the initial coarse disparity map. Statistical experimentation demonstrated a 3.14% reduction in endpoint error compared to state-of-the-art methods. With a processing time of approximately 30fps, satisfying the requirements of real-time operation applications. In order to validate the performance of the trained MMDENet in actual endoscopic images, we conduct both qualitative and quantitative analysis with 93.38% high precision, which holds great promise for applications in surgical navigation.
中科院2区
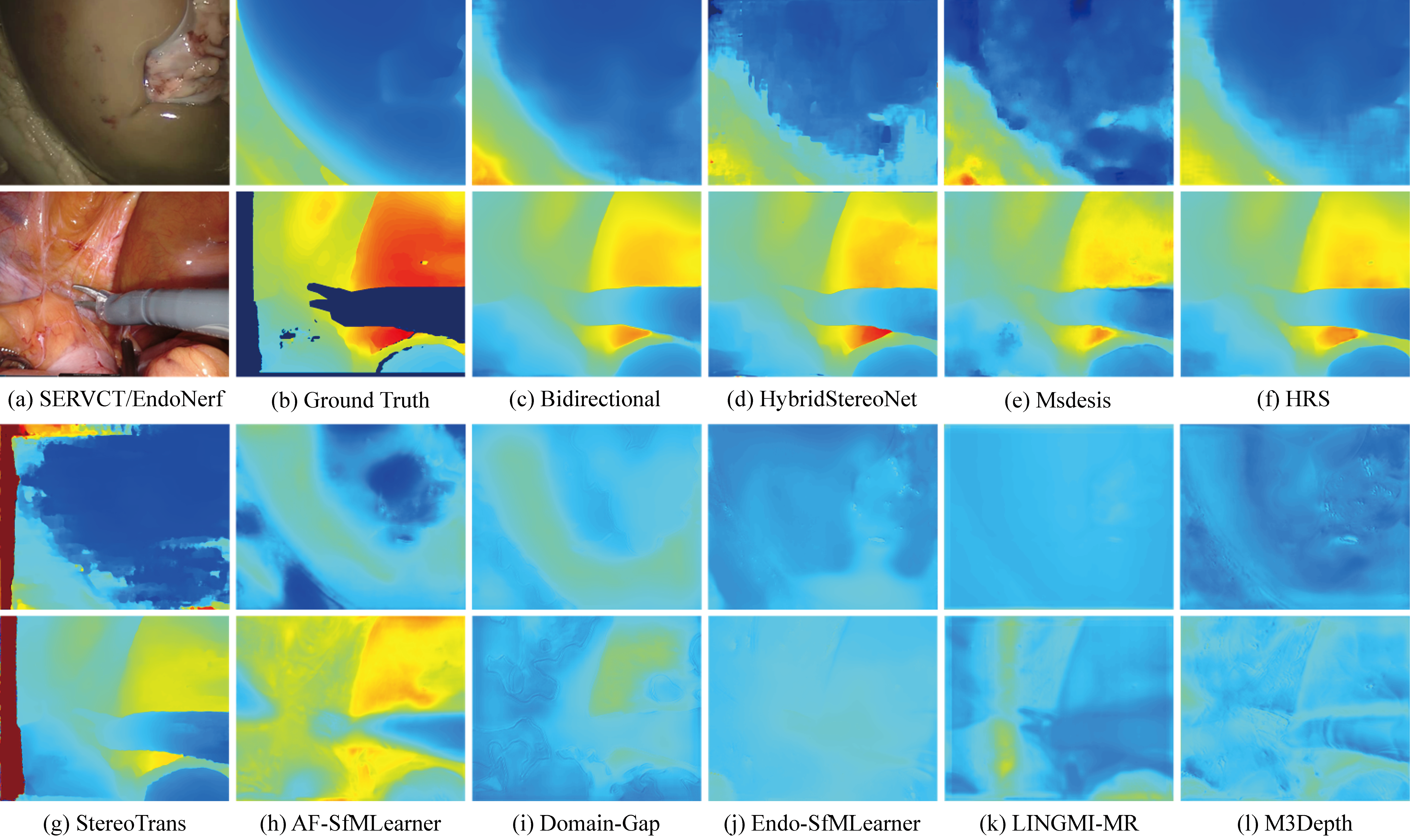
Deep learning for endoscopic depth estimation: A review
Xiongzhi Wang, Boyu Yang, Min Wei, Liangfa Liu, Jingang Zhang, Yunfeng Nie
Displays, 2025: 103086
- Reviews the application of deep learning in endoscopic depth estimation.
- Analyzes the characteristics of endoscopic datasets and metric selection.
- Compares supervised, self-supervised, and semi-supervised learning methods.
- Suggests future work: high-quality data and lightweight models for endoscopy.
[paper]
[abstract]
Depth estimation is a fundamental task in computer vision, crucial for applications such as endoscopic surgical navigation. This paper comprehensively reviews recent advancements in endoscopic depth estimation algorithms utilizing deep learning. We start by briefly describing the basic principles behind depth estimation and how depth maps can be generated from monocular and binocular cues. We then analyze the characteristics of the endoscopic dataset. Subsequently, we provide an overview of deep learning applications in endoscopic depth estimation, encompassing supervised, self-supervised, and semi-supervised learning methods. We examine each method’s principles, advantages, and disadvantages and their performance in practical applications. Additionally, we summarize the performance of current deep learning methods in endoscopic depth estimation and explore the importance of model robustness and generalization capabilities. Finally, we propose potential future research directions, such as exploring methods for collecting high-quality data or using simulated data to overcome current dataset limitations, and developing lightweight models to enhance real-time performance and robustness. This study aims to offer a comprehensive review for researchers in the field of endoscopic depth estimation, thereby fostering further development in this area.
中科院2区
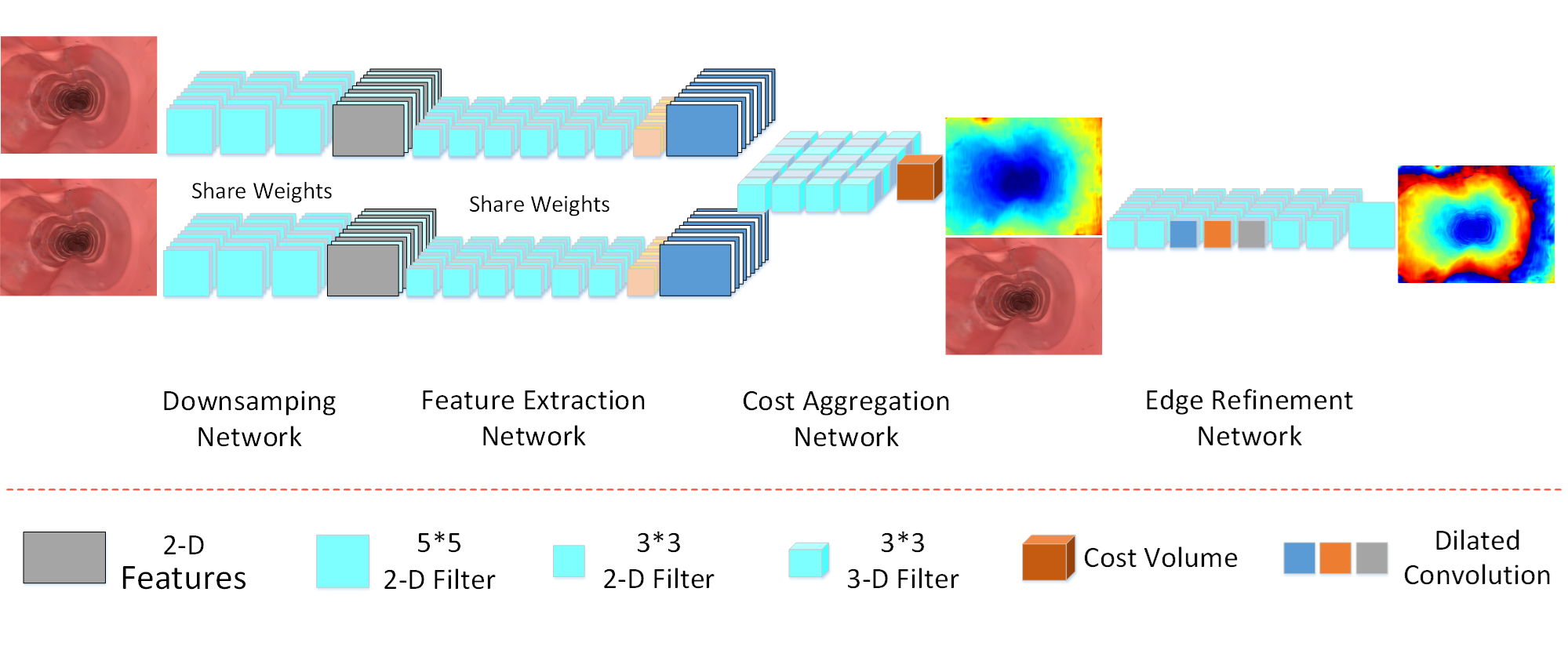
Deep convolutional network for stereo depth mapping in binocular endoscopy
Xiongzhi Wang, Yunfeng Nie, Shaoping Lu, Jingang Zhang
IEEE Access, 2020, 8: 73241–73249
- Generates binocular endoscopy datasets via 3D gastrointestinal simulation
- Proposes a 23-layer lightweight CNN for real-time stereo depth mapping
- Designs a scale-invariant loss function for endoscopic image characteristics
[paper]
[abstract]
Depth mapping from binocular endoscopy images plays an important role in stereoscopic surgical treatment. Owing to the development of deep convolutional neural networks (CNNs), binocular depth estimation models have achieved many exciting results in the felds of autonomous driving and machine vision. However, the application of these methods to endoscopic imaging is greatly limited by the fact that binocular endoscopic images not only are rare, but also have unsatisfying features such as no texture,
no ground truth, bad contrast, and high gloss. Aiming at solving the above-mentioned problems, we have built a precise gastrointestinal environment by the open-source software blender to simulate abundant binocular endoscopy data and proposed a 23-layer deep CNNs method to generate real-time stereo depth mapping. An effcient scale-invariant loss function is introduced in this paper to accommodate the characteristics of endoscope images, which improves the accuracy of achieved depth mapping results. Regarding the
considerable training data for typical CNNs, our method requires only a few images (960 * 720 resolution) at 45 frames per second on an NVIDIA GTX 1080 GPU module, then the depth mapping information is generated in real-time with satisfactory accuracy. The effectiveness of the developed method is validated by comparing with state-of-the-art methods on processing the same datasets, demonstrating a faster and more accurate performance than other model frames.
Deep convolutional network for stereo depth mapping in binocular endoscopy
**Xiongzhi Wang**, Yunfeng Nie, Shaoping Lu, Jingang Zhang
*IEEE Access*, 2020, 8: 73241–73249
Multi-scale, multi-dimensional binocular endoscopic image depth estimation network
**Xiongzhi Wang**, Yunfeng Nie, Wenqi Ren, Min Wei, Jingang Zhang
*Computers in Biology and Medicine*, 2023, 164: 107305
Deep learning for endoscopic depth estimation: A review
**Xiongzhi Wang**, Boyu Yang, Min Wei, Liangfa Liu, Jingang Zhang, Yunfeng Nie
*Displays*, 2025: 103086
Endo-E2E-GS: End-to-end 3D reconstruction of endoscopic scenes using Gaussian Splatting
**Xiongzhi Wang**, Boyu Yang, Min Wei, Yu Chen, Jingang Zhang, Yunfeng Nie
*Displays*, 2026: 103353
📚 Books & Book Chapters
Research and Exploration of Endoscopic Image Big Data Technology Serving Precision Medicine
Jingang Zhang, Liang Zong, Min Wei, Wenqi Ren, Xiongzhi Wang, Yanbo Dong, Kun Liu, Liangfa Liu, Shiming Yang
China’s e-Science Blue Book 2023, Springer Nature, 2024: 397–413
计算工程光学(Python版)
张金刚,聂云峰,付强,王雄智
西安电子科技大学出版社,2023.07
# 🎖 Honors and Awards
- *2021.10* Lorem ipsum dolor sit amet, consectetur adipiscing elit. Vivamus ornare aliquet ipsum, ac tempus justo dapibus sit amet.
- *2021.09* Lorem ipsum dolor sit amet, consectetur adipiscing elit. Vivamus ornare aliquet ipsum, ac tempus justo dapibus sit amet.
📖 Educations
- 2021.03 - 2026.06, Instrument Science and Technology, School of Aerospace Science and Technology, Xidian University, Xi’an, China.
- 2018.09 - 2021.03, Computer Technology, School of Computer Science and Technology, Xidian University, Xi’an, China.
- 2014.09 - 2018.06, Information and Technology Science, College of Science, China University of Petroleum (East China), Qingdao, China.
# 💬 Invited Talks
- *2021.06*, Lorem ipsum dolor sit amet, consectetur adipiscing elit. Vivamus ornare aliquet ipsum, ac tempus justo dapibus sit amet.
- *2021.03*, Lorem ipsum dolor sit amet, consectetur adipiscing elit. Vivamus ornare aliquet ipsum, ac tempus justo dapibus sit amet. \| [\[video\]](https://github.com/)
💻 Teaching
- Intelligent Image Processing with Python, Teaching Assistant - Fall 2021-2025
📄 Academic Service
- Npj Digital Surgery, Scientific Reports, Scientific Data




